NAME
r.pops.spread - A dynamic species distribution model for pest or pathogen spread in forest or agricultural ecosystemsKEYWORDS
raster, spread, model, disease, pestSYNOPSIS
r.pops.spread
r.pops.spread --helpr.pops.spread [-lms] host=name total_plants=name infected=name [output=name] [output_series=basename] [stddev=name] [stddev_series=basename] [probability=name] [outside_spores=name] [treatments=name[,name,...]] [treatment_year=integer[,integer,...]] wind=string [moisture_coefficient_file=name] [temperature_coefficient_file=name] [lethal_temperature=float] [lethal_month=integer] [temperature_file=name] start_time=integer end_time=integer seasonality=from,to step=string [reproductive_rate=float] [dispersal_kernel=string] [short_distance_scale=float] [long_distance_scale=float] [kappa=float] [percent_short_dispersal=float] [mortality_rate=float] [mortality_time_lag=integer] [mortality_series=basename] [random_seed=integer] [runs=integer] [nprocs=integer] [--overwrite] [--help] [--verbose] [--quiet] [--ui]
Flags:
- -l
- The output series as a single run only, not average
- The first run will be used for output instead of average
- -m
- Apply mortality
- After certain number of years, start removing dead hosts from the infected pool with a given rate
- -s
- Generate random seed (result is non-deterministic)
- Automatically generates random seed for random number generator (use when you don't want to provide the seed option)
- --overwrite
- Allow output files to overwrite existing files
- --help
- Print usage summary
- --verbose
- Verbose module output
- --quiet
- Quiet module output
- --ui
- Force launching GUI dialog
Parameters:
- host=name [required]
- Input host raster map
- Number of hosts per cell.
- total_plants=name [required]
- Input raster map of total plants
- Number of all plants per cell
- infected=name [required]
- Input raster map of initial infection
- Number of infected hosts per cell
- output=name
- Name for output raster map
- output_series=basename
- Basename for output series
- stddev=name
- Standard deviations
- stddev_series=basename
- Basename for output series of standard deviations
- probability=name
- Infection probability (in percent)
- outside_spores=name
- Output vector map of spores or pest units outside of modeled area
- treatments=name[,name,...]
- Raster map(s) of treatments (treated 1, otherwise 0)
- treatment_year=integer[,integer,...]
- Years when treatment rasters are applied
- wind=string [required]
- Prevailing wind direction
- NONE means that there is no wind
- Options: N, NE, E, SE, S, SW, W, NW, NONE
- Default: NONE
- moisture_coefficient_file=name
- Input file with one moisture coefficient map name per line
- Moisture coefficient
- temperature_coefficient_file=name
- Input file with one temperature coefficient map name per line
- Temperature coefficient
- lethal_temperature=float
- Temperature at which the pest or pathogen dies
- The temerature unit must be the same as for thetemerature raster map (typically degrees of Celsius)
- lethal_month=integer
- Month when the pest or patogen dies due to low temperature
- The temperature unit must be the same as for thetemperature raster map (typically degrees of Celsius)
- temperature_file=name
- Input file with one temperature raster map name per line
- The temperature should be in actual temperature units (typically degrees of Celsius)
- start_time=integer [required]
- Start year of the simulation
- The first day of the year will be used
- end_time=integer [required]
- End year of the simulation
- The last day of the year will be used
- seasonality=from,to [required]
- Seasonal spread (from,to)
- Spread limited to certain months (season), for example 5,9 for spread starting at the beginning of May and ending at the end of September
- Default: 1,12
- step=string [required]
- Simulation step
- How often the simulation computes new step
- Options: week, month
- week: Compute next simulation step each week
- month: Compute next simulation step each month
- reproductive_rate=float
- Number of spores or pest units produced by a single host
- Number of spores or pest units produced by a single host under optimal weather conditions
- Default: 4.4
- dispersal_kernel=string
- Type of dispersal kernel
- Options: cauchy, cauchy_mix
- Default: cauchy
- short_distance_scale=float
- Distance scale parameter for short range dispersal kernel
- Default: 20.57
- long_distance_scale=float
- Distance scale parameter for long range dispersal kernel
- kappa=float
- Strength of the wind direction in the von-mises distribution
- Default: 2
- percent_short_dispersal=float
- Percentage of short range dispersal
- What percentage of dispersal is short range versus long range
- Options: 0-1
- mortality_rate=float
- Mortality rate of infected hosts
- Percentage of infected hosts that die in a given year (hosts are removed from the infected pool)
- Options: 0-1
- mortality_time_lag=integer
- Time lag from infection until mortality can occur in years
- How many years it takes for an infected host to die (value 1 for hosts dying at the end of the first year)
- mortality_series=basename
- Basename for series of number of dead hosts
- Basename for output series of number of dead hosts (requires mortality to be activated)
- random_seed=integer
- Seed for random number generator
- The same seed can be used to obtain same results or random seed can be generated by other means.
- runs=integer
- Number of simulation runs
- The individual runs will obtain different seeds and will be averaged for the output
- nprocs=integer
- Number of threads for parallel computing
- Options: 1-
Table of contents
DESCRIPTION
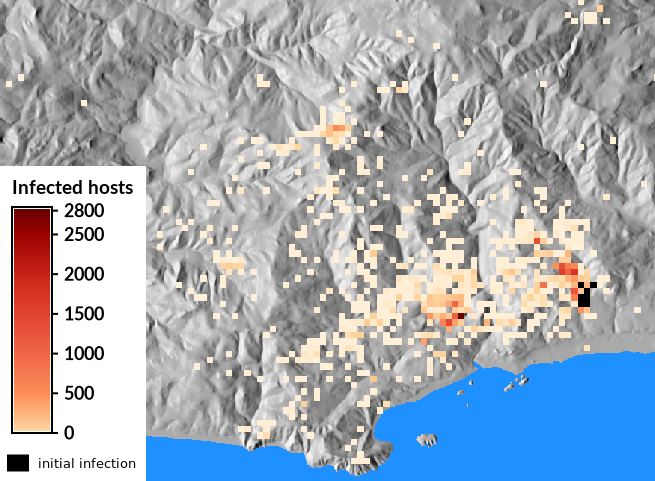
Module r.pops.spread is a dynamic species distribution model for pest or pathogen spread in forest or agricultural ecosystems. The model is process based meaning that it uses understanding of the effect of weather on reproduction and survival of the pest/pathogen in order to simulate spread of the pest/pathogen into the future.Module r.pops.spread is using Pest or Pathogen Spread library.
NOTES
- The directions of wind consider north (N) to be grid north, if your true north is different direction, you need to make an adjustment.
-
The module currently does not handle NULL (no data) as input, so you
need to change the NULLs to (most likely) zeros, for example:
r.null map=infection null=0.
EXAMPLES
Obtaining list of rasters
Use R script to create weather coefficients based on a defined polynomial.Example of creating file with list of input maps (unix-like command line):
g.list type=raster pattern="moisture_*" mapset=climate -m > moistures.txt g.list type=raster pattern="temperature_*" mapset=climate -m > temperatures.txt
g.list type=raster pattern="temperature_*" mapset=climate | sort -k2 -t_ -n > temperatures.txt
g.mapsets mapset=climate
Generating a constant coefficient
In case the moisture coefficient is not used, we can generate a constant raster map to be used as the coefficient:r.mapcalc "const_1 = 1"
NUM_LINES=`cat temperatures.txt | wc -l` echo const_1 > moistures.txt for LINE in `seq 2 $NUM_LINES`; do echo const_1 >> moistures.txt; done;
Creating treatments
To account for (vector) treatments partially covering host cells:# set resolution for treatments and convert to raster g.region res=10 -ap v.to.rast input=treatment output=treatment use=val # resample to lower resolution (match host map resolution) g.region align=host_map -p r.resamp.stats -w input=treatment output=treatment_resampled method=count # get maximum value, which is dependent on resolution # e.g. when resampling from 10m to 100m, max will be 100 (100 small cells in 1 big cell) r.info -r treatment_resampled # result will be 0 to 1 r.mapcalc "treatment_float = test_treatment_resampled / 100" # adjust host layer r.mapcalc "treated_host = host - host * treatment_float"
Running the model
Example of the run of the model (unix-like command line):
r.spread.pest host=host total_plants=all infected=infected_2005 \
moisture_coefficient_file=moistures.txt temperature_coefficient_file=temperatures.txt \
output=spread step=week start_time=2005 end_time=2010 \
reproductive_rate=4 dispersal_kernel=cauchy wind=NE random_seed=4
REFERENCES
- Ross K. Meentemeyer, Nik J. Cunniffe, Alex R. Cook, Joao A. N. Filipe, Richard D. Hunter, David M. Rizzo, and Christopher A. Gilligan 2011. Epidemiological modeling of invasion in heterogeneous landscapes: spread of sudden oak death in California (1990-2030). Ecosphere 2:art17. DOI: 10.1890/ES10-00192.1
- Tonini, Francesco, Douglas Shoemaker, Anna Petrasova, Brendan Harmon, Vaclav Petras, Richard C. Cobb, Helena Mitasova, and Ross K. Meentemeyer. Tangible geospatial modeling for collaborative solutions to invasive species management. Environmental Modelling & Software 92 (2017): 176-188. DOI: 10.1016/j.envsoft.2017.02.020
SEE ALSO
r.pops.spread on GitHubr.spread
AUTHORS
Francesco Tonini* (original R version)Zexi Chen* (C++ version)
Vaclav Petras* (parallelization, GRASS interface)
Anna Petrasova* (single species simulation)
* Center for Geospatial Analytics, NCSU
SOURCE CODE
Available at: r.pops.spread source code (history)
Main index | Raster index | Topics index | Keywords index | Graphical index | Full index
© 2003-2019 GRASS Development Team, GRASS GIS 7.4.5svn Reference Manual