
Note: A new GRASS GIS stable version has been released: GRASS GIS 7.8, available here.
Updated manual page: here
NAME
v.net.distance - Computes shortest distance via the network between the given sets of features.Finds the shortest paths from each 'from' point to the nearest 'to' feature and various information about this relation are uploaded to the attribute table.
KEYWORDS
vector, network, shortest pathSYNOPSIS
Flags:
- -g
- Use geodesic calculation for longitude-latitude locations
- -l
- Write each output path as one line, not as original input segments.
- --overwrite
- Allow output files to overwrite existing files
- --help
- Print usage summary
- --verbose
- Verbose module output
- --quiet
- Quiet module output
- --ui
- Force launching GUI dialog
Parameters:
- input=name [required]
- Name of input vector map
- Or data source for direct OGR access
- output=name [required]
- Name for output vector map
- arc_layer=string
- Arc layer
- Vector features can have category values in different layers. This number determines which layer to use. When used with direct OGR access this is the layer name.
- Default: 1
- arc_type=string[,string,...]
- Arc type
- Input feature type
- Options: line, boundary
- Default: line,boundary
- node_layer=string
- Node layer
- Vector features can have category values in different layers. This number determines which layer to use. When used with direct OGR access this is the layer name.
- Default: 2
- from_layer=string
- From layer number or name
- Vector features can have category values in different layers. This number determines which layer to use. When used with direct OGR access this is the layer name.
- Default: 1
- from_cats=range
- From category values
- Example: 1,3,7-9,13
- from_where=sql_query
- From WHERE conditions of SQL statement without 'where' keyword
- Example: income < 1000 and population >= 10000
- to_layer=string
- Layer number or name
- To layer number or name
- Default: 1
- to_type=string[,string,...]
- To feature type
- Options: point, line, boundary
- Default: point
- to_cats=range
- To category values
- Example: 1,3,7-9,13
- to_where=sql_query
- To WHERE conditions of SQL statement without 'where' keyword
- Example: income < 1000 and population >= 10000
- arc_column=name
- Arc forward/both direction(s) cost column (number)
- arc_backward_column=name
- Arc backward direction cost column (number)
- node_column=name
- Node cost column (number)
Table of contents
DESCRIPTION
v.net.distance finds the nearest element in set to for every point in set from.NOTES
These two sets are given by the respective layer, where and cats parameters. The type of to features is specified by to_type parameter. All from features are points. A table is linked to output map containing various information about the relation. More specifically, the table has three columns: cat, tcat and dist storing category of each from feature, category of the nearest to feature and the distance between them respectively.Furthemore, the output map contains the shortest path between each cat, tcat pair. Each path consists of several lines. If a line is on the shortest path from a point then the category of this point is assigned to the line. Note that every line may contain more than one category value since a single line may be on the shortest path for more than one from feature. And so the shortest paths can be easily obtained by querying lines with corresponding category number. Alternatively, unique paths can be created with the -l flag where each path will be a separate single line in the output.
The costs of arcs in forward and backward direction are specified by arc_column and arc_backward_column columns respectively. If arc_backward_column is not given, the same cost is used in both directions.
v.net.distance will not work if you are trying to find the nearest neighbors within a group of nodes, i.e. where to and from are the same set of nodes, as the closest node will be the node itself and the result will be zero-length paths. In order to find nearest neighbors within a group of nodes, you can either loop through each node as to and all other nodes as from or create a complete distance matrix with v.net.allpairs and select the lowest non-zero distance for each node.
EXAMPLES
Shortest path and distance between school and nearest hospital
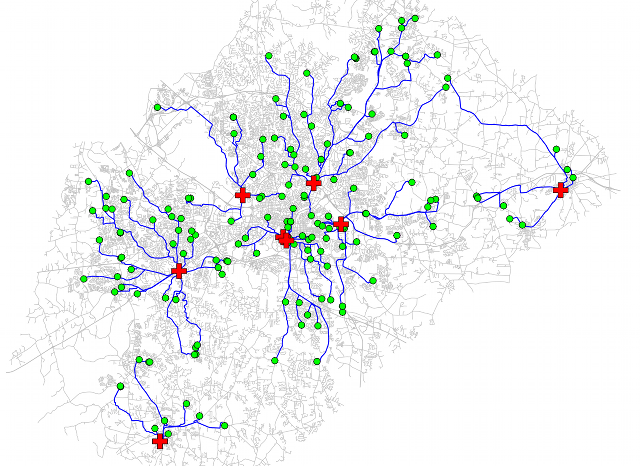
Find shortest path and distance from every school to the nearest hospital and show all paths.Streets are grey lines, schools are green circles, hospitals are red crosses, shortest paths are blue lines:
# connect schools to streets as layer 2
v.net input=streets_wake points=schools_wake output=streets_net1 \
operation=connect thresh=400 arc_layer=1 node_layer=2
# connect hospitals to streets as layer 3
v.net input=streets_net1 points=hospitals output=streets_net2 \
operation=connect thresh=400 arc_layer=1 node_layer=3
# inspect the result
v.category in=streets_net2 op=report
# shortest paths from schools (points in layer 2) to nearest hospitals (points in layer 3)
v.net.distance in=streets_net2 out=schools_to_hospitals flayer=2 to_layer=3
# visualization
g.region vector=streets_wake
d.mon wx0
d.vect streets_wake color=220:220:220
d.vect schools_wake color=green size=10
d.vect map=hospitals icon=basic/cross3 size=15 color=black fcolor=red
d.vect schools_to_hospitals
Distance between point source of pollution and sample points along streams
Example with streams of the NC sample data set.
# add coordinates of pollution point source of pollution as vector
pollution.txt:
634731.563206905|216390.501834892
v.in.ascii input=pollution.txt output=pollution
# add table to vector
v.db.addtable map=pollution
# add coordinates of sample points as vector
samples.txt:
634813.332814905|216333.590706166
634893.462007813|216273.763350851
634918.660011082|216254.949609689
v.in.ascii input=samples.txt output=samples
# add table to vector
v.db.addtable map=samples
# connect samples and pollution to streams
v.net -c input=streams points=samples output=streams_samples \
operation=connect node_layer=3 threshold=10 \
v.net -c input=streams_samples points=pollution
output=streams_samples_pollution operation=connect \
node_layer=4 threshold=10
# check vector layers
v.category input=streams_samples_pollution option=report
Layer/table: 1/streams_samples_pollution
type count min max
point 0 0 0
line 8562 40102 101351
boundary 0 0 0
centroid 0 0 0
area 0 0 0
face 0 0 0
kernel 0 0 0
all 8562 40102 101351
Layer: 3
type count min max
point 3 1 3
line 0 0 0
boundary 0 0 0
centroid 0 0 0
area 0 0 0
face 0 0 0
kernel 0 0 0
all 3 1 3
Layer: 4
type count min max
point 1 1 1
line 0 0 0
boundary 0 0 0
centroid 0 0 0
area 0 0 0
face 0 0 0
kernel 0 0 0
all 1 1 1
# calculate distance between sample points and pollution point source
v.net.distance input=streams_samples_pollution \
output=distance_samples_to_pollution from_layer=3 to_layer=4
# check results
v.report map=distance_samples_to_pollution@vnettest option=length
cat|tcat|dist|length
1|1|100.0|100.0
2|1|200.0|200.0
3|1|231.446|231.446
SEE ALSO
v.net.path, v.net.allpairs, v.net.distance, v.net.allocAUTHORS
Daniel Bundala, Google Summer of Code 2009, StudentWolf Bergenheim, Mentor
Markus Metz
Last changed: $Date$
SOURCE CODE
Available at: v.net.distance source code (history)
Note: A new GRASS GIS stable version has been released: GRASS GIS 7.8, available here.
Updated manual page: here
Main index | Vector index | Topics index | Keywords index | Graphical index | Full index
© 2003-2020 GRASS Development Team, GRASS GIS 7.6.2dev Reference Manual